- Blog
- Madden nfl 20 switch
- Hd forza horizon 4 wallpaper
- Lego pirates of the caribbean wii
- Hulu price increase
- Mcafee total protection vs internet security
- New game plus thief simulator
- Anime landscape wallpapers 1920 x 1080
- Twisted metal 4 rom ps1
- Spear girl nasuverse
- Omniplan pro 3
- Dymo stamps for mac download
- Man walking with little girl on moving dock
- Mp3 normalizer torrent
- Cellprofiler imge processing
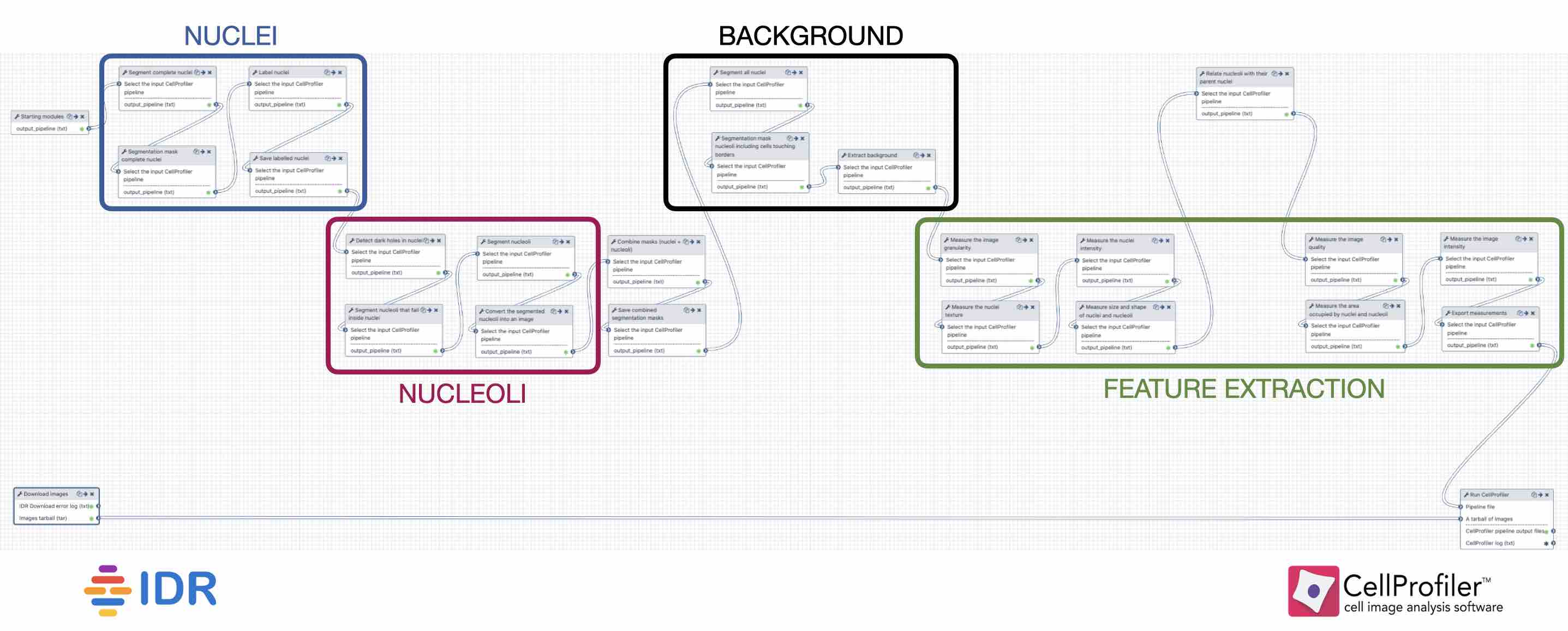
(E) Image processing done using Fiji’s MorphoLibJ plugin (macro code is presented in S1 Table). (C) CellProfiler 3.0 image processing modules used for HL60 cell nucleus segmentation. Both were compared to manually annotated ground truth using CellProfiler’s MeasureImageOverlap module. (B) Evaluation of CellProfiler 3.0 performance in comparison to the MorphoLibJ plugin in Fiji software. (A) Original 3D image of HL60 nuclei prior to analysis. The data set was generated using CytoPacq set up to simulate a Zeiss Axiovert S100 microscope (objective Zeiss 63×/1.40 Oil DIC) attached to confocal unit Atto CARV and CCD camera Micromax 1300-YHS. Synthetic images with 75% clustering probability and low SNR were chosen for analysis. Images are available from the Broad Bioimage Benchmark Collection ( ), as in Fig 2C of the main paper. S4 Fig: Segmentation steps for the analysis of synthetic images depicting HL60 cell line nuclei. 5 μ m) and provided by Javier Frias Aldeguer and Nicolas Rivron from Hubrecht Institute, Netherlands, and ground truth was annotated by Li Linfeng from MERLN Institute, Netherlands. Images were obtained using a PerkinElmer Ultraview VoX spinning disk microscope with a 63× oil immersion objective (distance between Z-slices = 0.
Cellprofiler imge processing manual#
(D) Ground truth obtained by manual annotation of each Z-slice using GIMP software. (C) CellProfiler 3.0 image processing modules used for stem cell nuclei. (A) Original 3D stem cell nuclei image prior to analysis. Images are available from the Broad Bioimage Benchmark Collection ( ), as in Fig 2B of the main paper. S3 Fig: Segmentation steps for the analysis of mouse trophoblast stem cell nuclei stained with Hoechst. 5 μ m) and provided by Javier Frias Aldeguer and Nicolas Rivron from Hubrecht Institute, Netherlands. Images were obtained using PerkinElmer Ultraview VoX spinning disk microscope with a 63× immersion objective (distance between Z-slices = 0. (C) CellProfiler 3.0 image processing modules used for blastocyst nuclei segmentation. (A) Original 3D image of blastocyst nuclei prior to analysis. Images are available from the Broad Bioimage Benchmark Collection ( ), as in Fig 2A of the main paper. S2 Fig: Segmentation steps for the analysis of mouse embryo blastocyst nuclei stained with Hoechst. Carpenter, Conceptualization, Funding acquisition, Project administration, Resources, Supervision, Writing – original draft, Writing – review & editing 1, * Caicedo, Software, Writing – review & editing, 1 and Anne E. Rafelski, Conceptualization, Methodology, Project administration, Resources, Supervision, Writing – review & editing, 4 Derek Thirstrup, Data curation, Methodology, Software, Validation, Writing – review & editing, 4 Winfried Wiegraebe, Conceptualization, Methodology, Resources, Supervision, 4 Shantanu Singh, Resources, Software, Writing – review & editing, 1 Tim Becker, Investigation, Software, Visualization, Writing – review & editing, 1 Juan C. Karhohs, Investigation, Methodology, Software, # 1 Minh Doan, Methodology, Software, Writing – original draft, Writing – review & editing, 1 Liya Ding, Data curation, Methodology, Software, Validation, Writing – review & editing, 4 Susanne M. Cimini, Conceptualization, Investigation, Methodology, Project administration, Software, Validation, Writing – review & editing, # 1 Kyle W. Claire McQuin, Conceptualization, Formal analysis, Software, Writing – original draft, Writing – review & editing, # 1 Allen Goodman, Conceptualization, Investigation, Software, # 1 Vasiliy Chernyshev, Data curation, Investigation, Methodology, Software, Validation, Visualization, Writing – review & editing, # 2, 3 Lee Kamentsky, Conceptualization, Software, # 1 Beth A.
